pymc.TruncatedNormal#
- class pymc.TruncatedNormal(name, *args, rng=None, dims=None, initval=None, observed=None, total_size=None, transform=UNSET, default_transform=UNSET, **kwargs)[source]#
Univariate truncated normal log-likelihood.
The pdf of this distribution is
\[f(x;\mu ,\sigma ,a,b)={\frac {\phi ({\frac {x-\mu }{\sigma }})}{ \sigma \left(\Phi ({\frac {b-\mu }{\sigma }})-\Phi ({\frac {a-\mu }{\sigma }})\right)}}\]Truncated normal distribution can be parameterized either in terms of precision or standard deviation. The link between the two parametrizations is given by
\[\tau = \dfrac{1}{\sigma^2}\](
Source code,png,hires.png,pdf)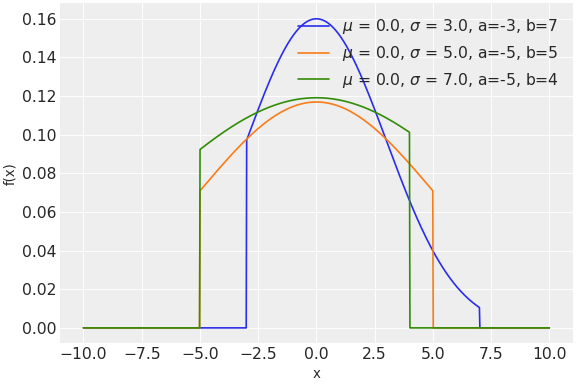
Support
\(x \in [a, b]\)
Mean
\(\mu +{\frac {\phi (\alpha )-\phi (\beta )}{Z}}\sigma\)
Variance
\(\sigma ^{2}\left[1+{\frac {\alpha \phi (\alpha )-\beta \phi (\beta )}{Z}}-\left({\frac {\phi (\alpha )-\phi (\beta )}{Z}}\right)^{2}\right]\)
- Parameters:
- mutensor_like of
float, default 0 Mean.
- sigmatensor_like of
float, optional Standard deviation (sigma > 0) (only required if tau is not specified). Defaults to 1 if neither sigma nor tau is specified.
- tautensor_like of
float, optional Precision (tau > 0) (only required if sigma is not specified).
- lowertensor_like of
float, default -numpy.inf Left bound.
- uppertensor_like of
float, defaultnumpy.inf Right bound.
- mutensor_like of
Examples
with pm.Model(): x = pm.TruncatedNormal('x', mu=0, sigma=10, lower=0) with pm.Model(): x = pm.TruncatedNormal('x', mu=0, sigma=10, upper=1) with pm.Model(): x = pm.TruncatedNormal('x', mu=0, sigma=10, lower=0, upper=1)
Methods
TruncatedNormal.dist([mu, sigma, tau, ...])Creates a tensor variable corresponding to the cls distribution.